
英文是纯手打的!论文原文的summarizing and paraphrasing。可能会出现难以避免的拼写错误和语法错误,若有发现欢迎评论指正!文章偏向于笔记,谨慎食用
目录
1. 心得
(1)Workshop啦,主要是标题装不下了
(2)很短的论文正文只有四页,可以直接导航到主模型图看我中文解释一分钟搞定
(3)附录有组织图片还有一堆表的,感兴趣的可以论文里面自取,这里没有附录
2. 论文逐段精读
2.1. Abstract
①Existing methods to fuse global image-level and cell-level: MLP or transformer
oncology n.[肿瘤] 肿瘤学 histology n.组织学 colorectal adj.结肠直肠的 gastric adj.胃的;胃部的
2.2. Introduction
①They comibine CNN and GNN on training
2.3. Integrating CNN with GNN
①Overall framework:
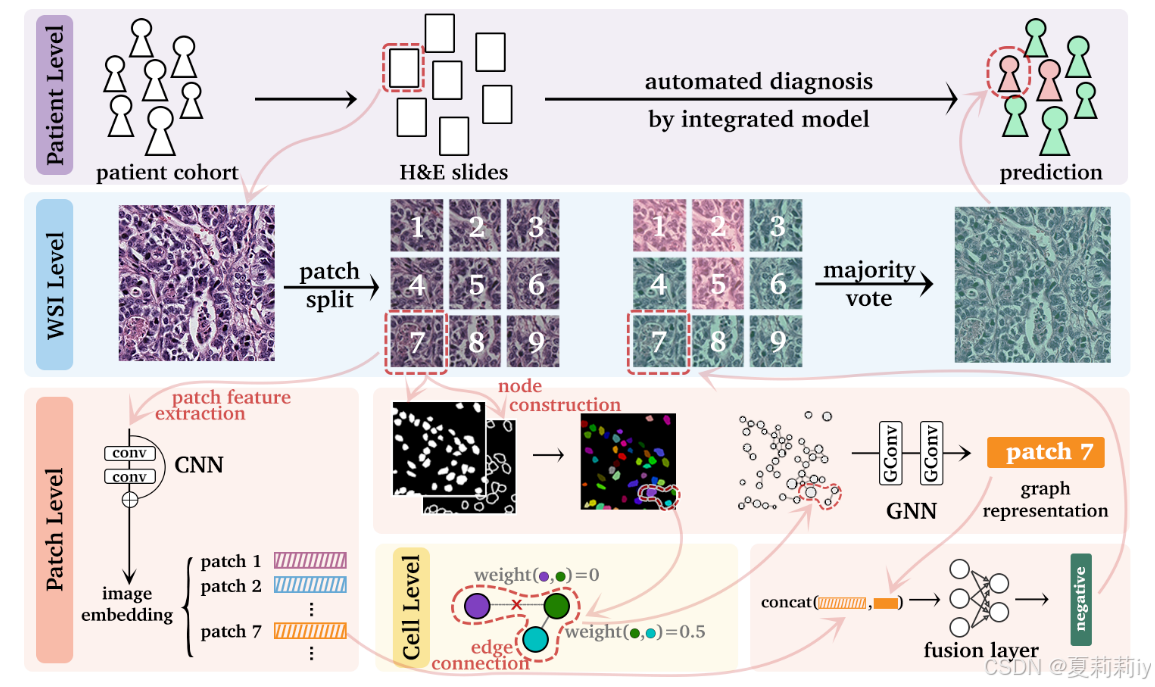
(我不是做这个方向的,我猜测,第一行:一个人的输入是一个组织切片,输出是是否患病?第二行是一个切片分成很多patch,然后第三行每个patch经过CNN提取特征,右边是每个patch的每个核一起全部变成一个图然后用GNN来卷,和CNN一样,GNN也是每个patch得到一个图的图级表示。最后把CNN和GNN每个patch的特征concat在一起就接MLP了。感觉是个很简单的模型,毕竟也比较早期的,写得还是很易于理解的没有弯弯绕绕)
②For stained histology whole-slide images (WSIs) with 224*224 resolution, they split each of them into non-overlaped patches
③Fusion strategy:
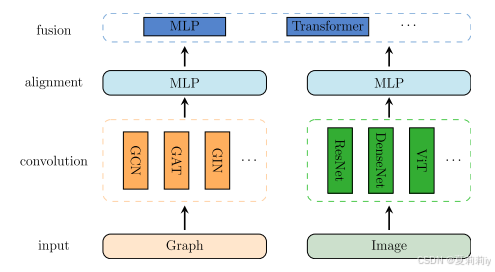
④Each patch can be regarded as a cell graph , where
denotes node set,
is edge set, each nuclei region in graph is a node
⑤Node features are extracted by CA2.5Net
⑥Edge is constructed by pair-wise Euclidean distance between nuclei centroids:
where denotes Euclidean distance,
denotes interaction between 2 cells. Adjacency matrix
⑦Hidden representation updating:
⑧Alignment of CNN such as DenseNet:
or ResNet:
⑨Combine output of CNN and GNN
by MLP:
where ,
⑩Combine output of CNN and GNN
by Transformer:
2.4. Experiments and Results
①Datasets: CR-MSI (binary microsatellite instability (MSI) status classification), STAD-MSI (binary microsatellite instability (MSI) status classification), and GIST-PDL1 (binary Programmed Death-Ligand 1 (PD-L1) status binary classification)
②CNN backbones: MOBILENETV3, DENSENET, and RESNET
③Performance table:
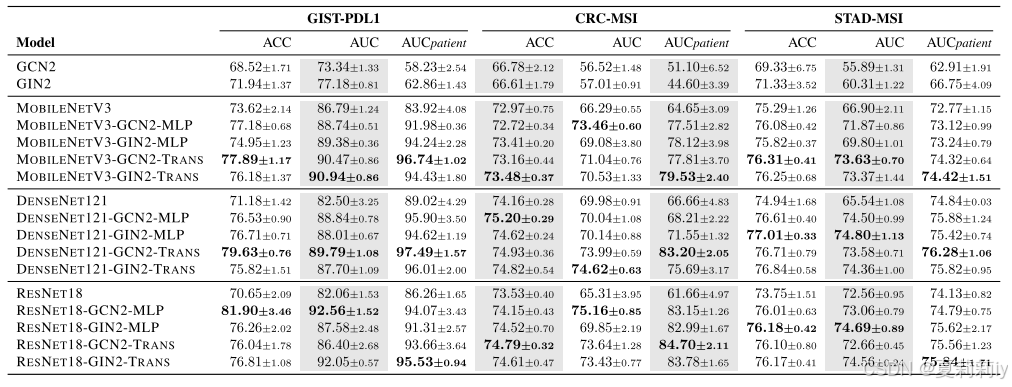
2.5. Discussion
~
3. Reference
Shen, Y. et al. (2022) How Graph Neural Networks Enhance Convolutional Neural Networks Towards Mining the Topological Structures from Histology, ICML Workshop on Computational Biology. Baltimore, Maryland, USA.
























 5657
5657

 被折叠的 条评论
为什么被折叠?
被折叠的 条评论
为什么被折叠?








