获取更好阅读体验:解决99%的GEO芯片平台探针基因注释问题!
1. 前言
上一篇文章 听说你的使用getGEO经常有网络问题?一行代码解决下载GEO表达谱和临床数据 解决了GEO表达谱下载的问题。
然而,从众多客户的需求来看,大家对于后续的探针注释似乎觉得更难。
那么,这次我就教大家如何进行探针的基因注释,基本可以涵盖 99% 平台的情况。
B站已经同步讲解 本篇内容,复制BV号【BV1ifDoY4E1b】至B站可直接观看。
本教程所有代码、资料已打包,公众号【生信摆渡】后台回复【download_GEO】即可获取。
2. GPL文件的下载和读入
虽然有一些自动注释的R包,但大家还是要学习如何从GPL文件自己构建注释表的,毕竟R包更新是不及时的。
以GPL570为例。
首先看网页,包含了头部信息 Data table header descriptions 和一部分的表格 Data table。
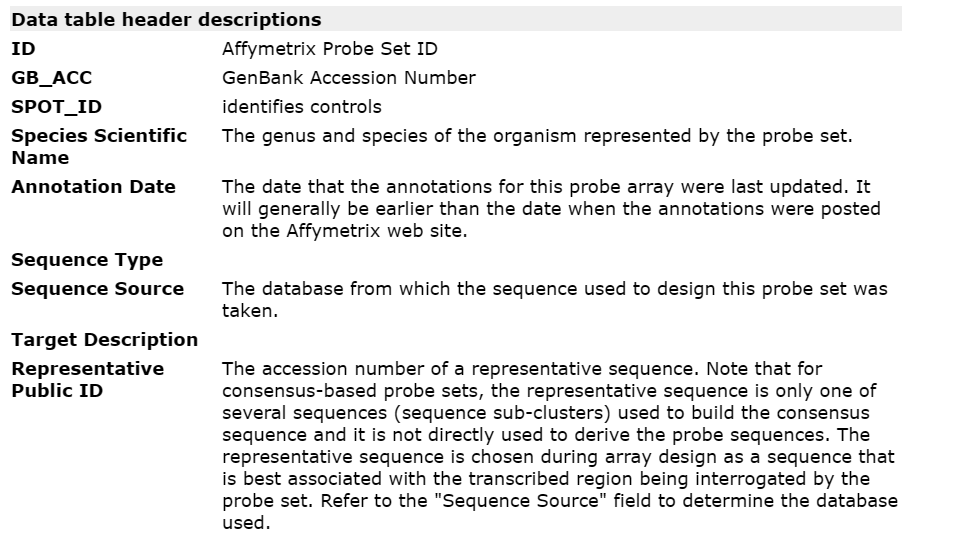

头部信息包含了注释表格每一列的解释。在GPL文件中以#标识符开头,因此在读入时需要跳过。
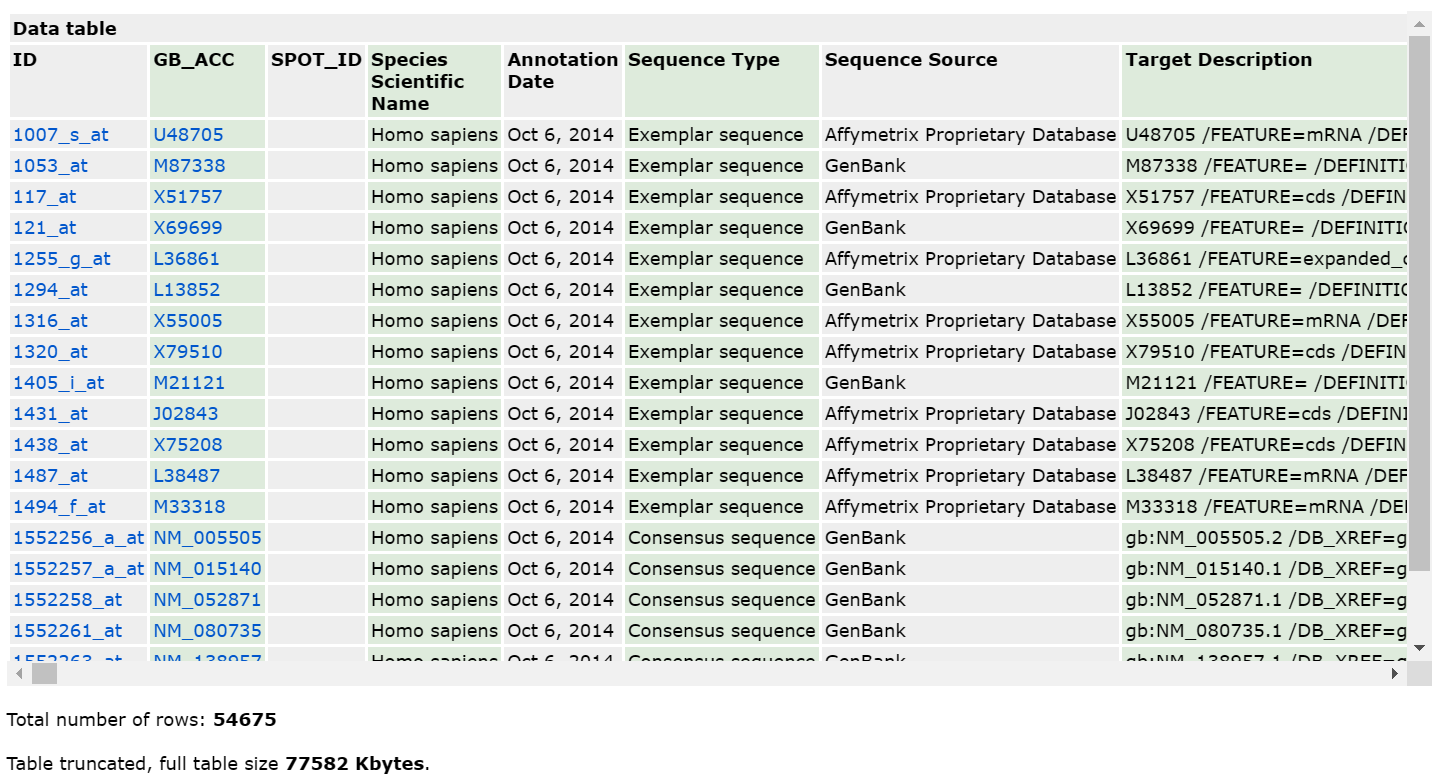
接着展示注释表格的行数和文件大小。
再往下是下载按钮。不过有的平台不提供下载按钮,只能在线查看,而且打开后浏览器就卡住了。
这种情况可以将在线查看的HTML网页下载下来,然后进行读取处理也是可以的,下面会详细介绍。
读取:
- 首先安装我编写的工具包
remotes::install_github("JiahaoWongg/bioinfobaidu.utils")
- 下载下来后进行读取
read_GPL <- function(GPL_file){
nskip = sum(subString(readr::read_lines(GPL_file, n_max = 100), 1) == "#")
anno_tb = data.table::fread(GPL_file, skip = nskip)
return(anno_tb)
}
GPL_file = "GPL570.txt"
raw_tb = read_GPL(GPL_file)
hd(raw_tb)
# dim: 54675 × 16
# ID GB_ACC SPOT_ID Species Scientific Name Annotation Date
# 1 1007_s_at U48705 Homo sapiens Oct 6, 2014
# 2 1053_at M87338 Homo sapiens Oct 6, 2014
# 3 117_at X51757 Homo sapiens Oct 6, 2014
# 4 121_at X69699 Homo sapiens Oct 6, 2014
# 5 1255_g_at L36861 Homo sapiens Oct 6, 2014
构建探针注释数据库
- 提取注释信息。
- 去除一探针对应多个基因的情况,只取第一个基因。
- 统一将没有对应基因的探针注释为空
""
anno_tb = data.frame(ID = raw_tb$ID, SYMBOL = raw_tb$`Gene Symbol`)
anno_tb$SYMBOL = subString(anno_tb$SYMBOL, 1, "[ /|]")
anno_tb$SYMBOL[anno_tb$SYMBOL == "NA"] = ""
anno_tb$SYMBOL[anno_tb$SYMBOL == "---"] = ""
hd(anno_tb)
# dim: 54675 × 2
# ID SYMBOL
# 1 1007_s_at DDR1
# 2 1053_at RFC2
# 3 117_at HSPA6
# 4 121_at PAX8
# 5 1255_g_at GUCA1A
这样注释文件就准备好了,为了方便使用,可以将每一个GPL注释数据库保存在一个列表里,并保存为RData,需要用时载入就行,这就构建了探针注释数据库了。
obj_file = "./anno_obj.RData"
if(!file.exists(obj_file)){
anno_obj = list()
} else {
anno_obj = get(load(obj_file))
}
anno_obj[[GPL]] = anno_tb
save(anno_obj, file = obj_file)
进行注释
首先需要下载数据集的表达矩阵和样本信息。
使用之前的文章听说你的使用getGEO经常有网络问题?一行代码解决下载GEO表达谱和临床数据 的getGEO2函数可以一键下载。
并且这一次对getGEO2也进行更新了,第一点是自动检测和执行是否需要进行log转化,第二点是使用limma进行样本间标准化。
所以需要后台获取最新代码,当然,这次的代码也整合了新版本的getGEO2。
obj = getGEO2("GSE23586")
# INFO [2024-11-09 16:10:40] Querying dataset: GSE23586 ...
# Try time 1 ... succeed.
# - GPL: GPL570
# - Found 6 samples, 38 metas.
# - Writting sample_anno to 00_GEO_data/GSE23586_sample_anno.xlsx.
# INFO [2024-11-09 16:10:42] Execute log(expM + 1) ...
# INFO [2024-11-09 16:10:43] Normalize between arrays ...
# - Successed, file save to 00_GEO_data/GSE23586.RData.
GPL = obj$GPL
expM = obj$expM
- 之后将原始表达矩阵的行名根据注释表替换为基因名就行了。
- 可能存在多探针对应同一个基因的情况,可以选择的去重方法可以选择取平均
mean或直接删掉重复的remove。建议取平均值,虽然计算要慢得多。如果很慢的话选择直接去除也是可以的。
anno_GEO <- function(expM, anno_tb, method = "mean"){
# check match
n_not_match = sum(! rownames(expM) %in% anno_tb$ID)
if(n_not_match > 0){
cat2("- Warnnig:", paste0(n_not_match, "/", nrow(expM)), "porbes fail to matched!\n")
} else {
cat2("- All porbes matched!\n")
}
# check SYMBOL
gene_name = anno_tb$SYMBOL[match(rownames(expM), anno_tb$ID)]
n_no_symbol = sum(gene_name == "", na.rm = TRUE)
if(n_no_symbol > 0){
cat2("- Warnning:", paste0(n_no_symbol, "/", nrow(expM)), "probes fail to annotated!\n")
} else {
cat2("- All porbes annotated!\n")
}
expM = data.frame(gene_name, expM)
expM2 = expM[expM$gene_name != "", ]
report(paste("Removing duplicated genes by method:", method, "..."))
if(method == "mean"){
expM3 = aggregate(expM2[, -1], by = list(factor(expM2[, 1])), mean0)
rownames(expM3) = expM3[, 1]
expM3[, 1] = NULL
} else if(method == "remove"){
expM3 = expM2[!duplicated(expM2[, 1]), ]
rownames(expM3) = expM3[, 1]
expM3[, 1] = NULL
}
return(expM3)
}
anno_list = get(load(anno_file))
if(GPL %in% names(anno_list)){
report("Use local annotation file ...")
anno_tb = anno_list[[GPL]]
expM_anno = anno_GEO(expM, anno_tb)
}
# INFO [2024-11-09 16:15:38] Use local annotation file ...
# - All porbes matched!
# - All porbes annotated!
# INFO [2024-11-09 16:15:38] Removing duplicated genes by method: mean ...
- 并提示有多少探针找不到注释信息或者注释信息为空。
- 至此,探针注释就完成了
hd(expM_anno)
# dim: 22881 × 6
# GSM578529 GSM578530 GSM578531 GSM578532 GSM578533
# A1BG 7.022058 7.809902 6.896205 7.037151 7.071827
# A1BG-AS1 7.168314 5.108887 6.941011 5.997464 4.591907
# A1CF 6.182022 6.001592 6.112716 6.823624 5.634210
# A2M 8.910845 9.665579 9.145603 9.548700 8.718978
# A2M-AS1 6.525060 7.602097 7.376438 7.080512 6.642209
R包自动注释
自动注释就是曾老师编写的AnnoProbe包。
我进行了一点的修改,就是前面所说的一探针对应多给予你的情况,这里貌似没有处理。
然后将注释数据库整理成统一的格式,方便统一调用。
anno_GEO_AnnoProbe <- function(expM, GPL, method = "mean"){
# idmap包可以被 AnnoProbe 包替代
# if(sum(c("idmap1", "idmap2", "idmap3") %in% data.frame(installed.packages())$Package) != 3){
# cat2("Please install annotation packages 'idmap1' 'idmap2' 'idmap3' from github.\n")
# stop()
# }
# loadp(idmap1, idmap2, idmap3)
# anno_tb1 = try(suppressMessages(getIDs(GPL)), silent = TRUE)
# anno_tb2 = try(suppressMessages(get_soft_IDs(GPL)), silent = TRUE)
# anno_tb3 = try(suppressMessages(get_pipe_IDs(GPL)), silent = TRUE)
# if(class(anno_tb1) == "data.frame"){
# anno_tb = anno_tb1
# } else if(class(anno_tb2) == "data.frame"){
# anno_tb = anno_tb2
# } else if(class(anno_tb3) == "data.frame"){
# anno_tb = anno_tb3
# } else {
# cat2("Can't find annotation table, please DIY one.\n")
# stop()
# }
if(! "AnnoProbe" %in% data.frame(installed.packages())$Package){
cat2("Please install annotation packages 'AnnoProbe' from github.\n")
cat2("\tdevtools::install_github(\"jmzeng1314/AnnoProbe\").\n")
stop()
}
report("Use AnnoProbe annotation file ...")
loadp(AnnoProbe)
idmap <- function (gpl = "GPL570", type = "bioc", mirror = "tercent"){
gpl = toupper(gpl)
gpl_anno = paste(gpl, c("bioc", "soft", "pipe"), sep = "_")
if (mirror == "tercent") {
up = "http://49.235.27.111"
}
if (!checkGPL(gpl)) {
stop("This platform is not in our list, please use our shinyAPP to custom annotate your probe sequences, or ask us to process and then update the R package!")
}
else {
tryCatch(utils::data("exists_anno_list", package = "AnnoProbe"))
gpl_anno = gpl_anno[gpl_anno %in% exists_anno_list]
if (T) {
tpf = paste0(paste(gpl, type, sep = "_"), ".rda")
down = paste0("/GEOmirror/GPL/", tpf)
download.file(paste0(up, down), tpf, mode = "wb", quiet = TRUE)
load(tpf)
return(get(paste(gpl, type, sep = "_")))
}
else {
stop("We have that platform, but just offer other type of annotaion.")
}
}
}
anno_tb = try(suppressMessages(idmap(GPL, type = "pipe")), silent = TRUE)
if(class(anno_tb) == "try-error"){
cat2("- Can't find annotation table in AnnoProbe.\n")
stop()
}
colnames(anno_tb) = c("ID", "SYMBOL")
anno_tb = anno_tb[!duplicated(anno_tb$ID), ]
anno_tb$SYMBOL = subString(anno_tb$SYMBOL, 1, "[ /|]")
anno_tb$SYMBOL[anno_tb$SYMBOL == "NA"] = ""
anno_tb$SYMBOL[anno_tb$SYMBOL == "---"] = ""
expM_anno = anno_GEO(expM, anno_tb)
return(expM_anno)
}
我们使用数据集GSE106090测试:
GSE = "GSE106090"
obj = getGEO2(GSE, out_dir = GSE)
# INFO [2024-11-09 16:22:56] Querying dataset: GSE106090 ...
# Try time 1 ... succeed.
# - GPL: GPL21827
# - Found 18 samples, 44 metas.
# - Writting sample_anno to GSE106090/GSE106090_sample_anno.xlsx.
# INFO [2024-11-09 16:22:59] Normalize between arrays ...
# - Successed, file save to GSE106090/GSE106090.RData.
GPL = obj$GPL
expM = obj$expM
expM_anno = anno_GEO_AnnoProbe(expM, GPL)
# INFO [2024-11-09 16:23:00] Use AnnoProbe annotation file ...
# - Warnnig: 16541/61046 porbes fail to matched!
# - All porbes annotated!
# INFO [2024-11-09 16:23:01] Removing duplicated genes by method: mean ...
hd(expM_anno)
# dim: 31119 × 18
# GSM2829419 GSM2829420 GSM2829421 GSM2829422 GSM2829423
# A1BG 5.057959 5.030532 4.999752 5.199815 4.931398
# A2M-AS1 8.921258 9.073332 9.113745 9.010616 9.206460
# A2ML1 5.405560 5.161947 6.053355 5.855274 6.034109
# A2ML1-AS2 2.517003 2.352829 2.348781 2.351604 3.349039
# A2MP1 3.413715 3.385146 3.259971 3.766047 3.531293
也是非常滴好用。
在线获取注释
后来,我又发现,有些平台有提供在线的GPLxxx.annot.gz文件,内容和下载GPLxxx.txt文件内容是一样的(头信息外)。所以在自动注释不能用且还没有制作本地注释文件的时候,可以尝试一下。
感觉提供这个文件的平台并不是太多,但不如让代码去试试~
至于为什么GPL网页注释文件那个下载按钮只能点击而没有的链接,我也是不明白,有固定链接的话就不用手动点击了。
同样,编写一个函数进行注释:
anno_GEO_online <- function(expM, GPL, out_dir = "annot_dir", symbol_name = NULL){
mkdir(out_dir)
GPL_num = gsub("GPL", "", GPL)
master_string = ifelse(nchar(GPL_num) < 4, "", subString(GPL_num, 1:(nchar(GPL_num) - 3)))
master_dir = paste0("GPL", master_string, "nnn")
url_prefix = "https://ftp.ncbi.nlm.nih.gov/geo/platforms"
soft_url = paste0(url_prefix, "/", master_dir, "/", GPL, "/annot/", GPL, ".annot.gz")
soft_file = paste0(out_dir, "/", GPL, ".annot.gz")
report("Download online annotation file ...")
options(timeout = 100)
dl_res = try(download.file(soft_url, destfile = soft_file, quiet = TRUE), silent = TRUE)
if(class(dl_res) == "try-error"){
cat2("Can't find online annotation table. Please download annotation table manually:\n")
cat2(paste0("\thttps://www.ncbi.nlm.nih.gov/geo/query/acc.cgi?acc=", GPL, "\n"))
stop()
}
anno_tb = suppressWarnings(data.table::fread(soft_file, data.table = FALSE))
if(is.null(symbol_name)){
SYMBOL_col_names = c("Gene symbol", "GENE_SYMBOL", "GeneSymbol", "Symbol", "SYMBOL", "gene_assignment")
symbol_name = SYMBOL_col_names[which(SYMBOL_col_names %in% colnames(anno_tb))]
if(length(symbol_name) == 0){
cat2("Can't find right 'symbol' column name. If exists exactly, please specific it to 'symbol_name'\n")
stop()
}
}
anno_tb = anno_tb[, c("ID", symbol_name)]
colnames(anno_tb) = c("ID", "SYMBOL")
anno_tb$SYMBOL = subString(anno_tb$SYMBOL, 1, "[ /|]")
anno_tb$SYMBOL[anno_tb$SYMBOL == "NA"] = ""
expM_anno = anno_GEO(expM, anno_tb)
return(expM_anno)
}
我们还是使用数据集GSE23586进行演示:
GSE = "GSE23586"
obj = download_GEO(GSE, out_dir = "GSE23586", anno_file = anno_file)
# INFO [2024-11-09 16:31:44] Querying dataset: GSE23586 ...
# Try time 1 ... succeed.
# - GPL: GPL570
# - Found 6 samples, 38 metas.
# - Writting sample_anno to GSE23586/GSE23586_sample_anno.xlsx.
# INFO [2024-11-09 16:31:46] Execute log(expM + 1) ...
# INFO [2024-11-09 16:31:46] Normalize between arrays ...
# - Successed, file save to GSE23586/GSE23586.RData.
GPL = obj$GPL
expM = obj$expM
expM_anno = anno_GEO_online(expM, GPL)
# INFO [2024-11-09 16:31:48] Download online annotation file ...
# - All porbes matched!
# - Warnning: 9557/54675 probes fail to annotated!
# INFO [2024-11-09 16:31:54] Removing duplicated genes by method: mean ...
hd(expM_anno)
# dim: 21754 × 6
# GSM578529 GSM578530 GSM578531 GSM578532 GSM578533
# A1BG 7.022058 7.809902 6.896205 7.037151 7.071827
# A1BG-AS1 7.168314 5.108887 6.941011 5.997464 4.591907
# A1CF 6.182022 6.001592 6.112716 6.823624 5.634210
# A2M 8.910845 9.665579 9.145603 9.548700 8.718978
# A2M-AS1 6.525060 7.602097 7.376438 7.080512 6.642209
也肥肠滴好用~
整合代码
不用担心代码很多,我最终把这三种注释方式整合为一个函数download_GEO了,包含表达谱的下载和注释,同时输出样本注释文件为csv格式。
如果你能正常使用xlsx包的话,还会输出xlsx格式的文件,好处是表格的列宽是自适应的,不需要你再因为字挡住而拖来拖去的了(能用代码解决绝不用手。。。
最后,再示范一下,非常简单:
GSE = "GSE23586"
obj = download_GEO(GSE, anno_file = anno_file)
# Step1: Download GEO data ...
# INFO [2024-11-09 16:39:44] Querying dataset: GSE23586 ...
# Try time 1 ... succeed.
# - GPL: GPL570
# - Found 6 samples, 38 metas.
# - Writting sample_anno to 00_GEO_data/GSE23586_sample_anno.xlsx.
# INFO [2024-11-09 16:39:47] Execute log(expM + 1) ...
# INFO [2024-11-09 16:39:47] Normalize between arrays ...
# - Successed, file save to 00_GEO_data/GSE23586.RData.
# Step2: Annotate probe ...
# INFO [2024-11-09 16:39:49] Use local annotation file ...
# - All porbes matched!
# - All porbes annotated!
# INFO [2024-11-09 16:39:49] Removing duplicated genes by method: mean ...
# INFO [2024-11-09 16:39:52] Done.
names(obj)
# [1] "GPL" "expM" "sample_anno"
- 结果就是一个列表,包含GPL索引、注释后的表达谱和样本注释信息。
- 其中
anno_file是我整理的注释文件,提供的压缩包里有。当然不提供这个文件也可以,就要碰碰运气看另外两种自动注释的方法能不能成功了。
这个数据集是可以的。
obj = download_GEO(GSE)
我相信,应该能够解决99%的注释问题,本地注释文件和代码我会一直更新,公众号关键词回复的链接也会同步更新。
学习更多生信技巧,请持续关注工种浩【生信摆渡】。
























 9005
9005

 被折叠的 条评论
为什么被折叠?
被折叠的 条评论
为什么被折叠?








